No products in the cart.
Bestsellers
Product categories
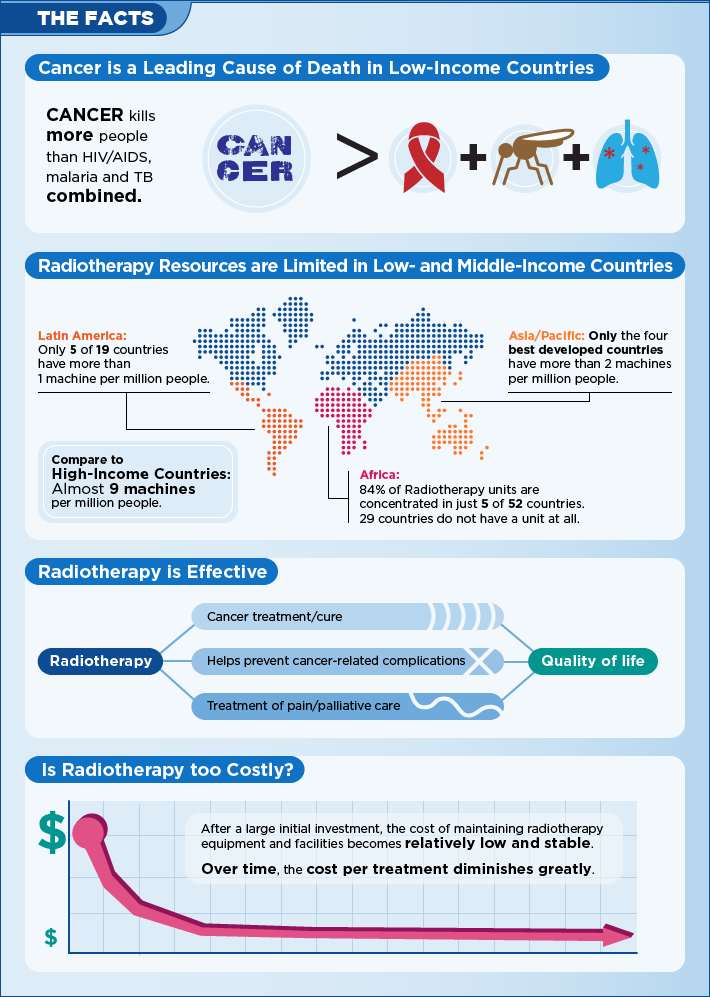
Men with Erectile Dysfunction in Australia
- Men with ED in percents
- Men with ED in percents
What Is It Like Living With Erectile Dysfunction In Australia
One thing that can seriously hurt a man's self-esteem is erectile dysfunction. This is a rather common problem for many men around the world and in Australia. In fact, around 1 million men are affected by erectile dysfunction in Australia. That makes it a rather common problem yeah it is not openly discussed. Many people around you are probably suffering from erectile dysfunction and you have no idea. Probably their Partners don't even have any idea because they may not even be honest with them. It can be so embarrassing that many people choose to keep it a secret. Here is why that isn't a good idea and it's better to get it out in the open so that you can treat it and move past it.
What is erectile dysfunction
If more men spoke up about their erectile dysfunction then perhaps there would be less stigmatism surrounding it. Instead, men choose to keep their shortcomings hidden and locked away behind closed doors. This is not anything to be embarrassed about but rather it's a natural instinct to protect oneself. The reality is that today we have many easy solutions for erectile dysfunction and it's nothing to be ashamed of. Growing old is a part of life and everyone grows old. Not a single person and escapes the grip of time and the effects it has on the human body. There are many solutions to erectile dysfunction with the most common being in pill form called Viagra.
What is Viagra
Viagra is an erectile dysfunction medication that is commonly used to help men regain an erection. When men are no longer able to get an erection they must turn to an artificial booster. It is nothing to be ashamed of if you require a medication in order to get an erection. Viagra contains an active ingredient called sildenafil which sends increased blood flow to the penis allowing for an erection when a man becomes sexually stimulated. This happens after a dose of Viagra. Viagra comes in 25mg, 50mg, and 100mg and is typically taken and doses of 25 to 50 mg at a time. Some people may even choose to take a smaller dose to start with and work their way up from there. Viagra is a small blue oblong pill that comes in 25 mg at the smallest dose.
What are the side effects
There are some side effects associated with taking erectile dysfunction medication but they are relatively minor. You may experience blurred vision, headaches, nausea, or muscle or back pain. These are all typical things associated with virtually any erectile dysfunction medication so you must weigh the cost was the benefit. In most cases, men don't experience any side effects at all and receive a healthy boost to their natural blood flow. This is a good way to get an erection back when you have been unable to get one. It is a simple matter of carrying a pill with you perhaps in your wallet so that you are ready for when the moment is right. One dose of Viagra will last between 30 minutes to 5 hours but typically last somewhere in between.
Who suffers from ED
Erectile dysfunction is probably more common among men than you realize. Since men all get older just at different rates they probably have experienced erectile dysfunction and if kept it to themselves. The best way to treat erectile dysfunction is by being honest with your doctor and explaining all the symptoms you are experiencing. They need all the details in order to make an accurate assessment and diagnosis. Once your doctor has done their exams and is ready to provide a diagnosis they will most likely prescribe a form of Viagra. From there they will simply write a prescription which can be filled at any pharmacy in Australia. Sometimes you can even find Viagra or generic versions from online websites and online pharmacies.
Check with your local laws about whether you can have them shipped to you.
How ED can be resolved
Don't be discouraged if you are experiencing erectile dysfunction because it is a common issue that is easily resolved. Most men can simply take and erectile dysfunction medication and regain the use of their penis. Just don’t make the mistake of keeping your problem secret and avoiding the doctor.